- Quick and Easy Workflow
- Bias-Free Nucleic Acid Extraction
- Inhibitor-Free DNA/RNA
- Certified Low Bioburden
- High-Throughput Automation
BIND, WASH, ELUTE
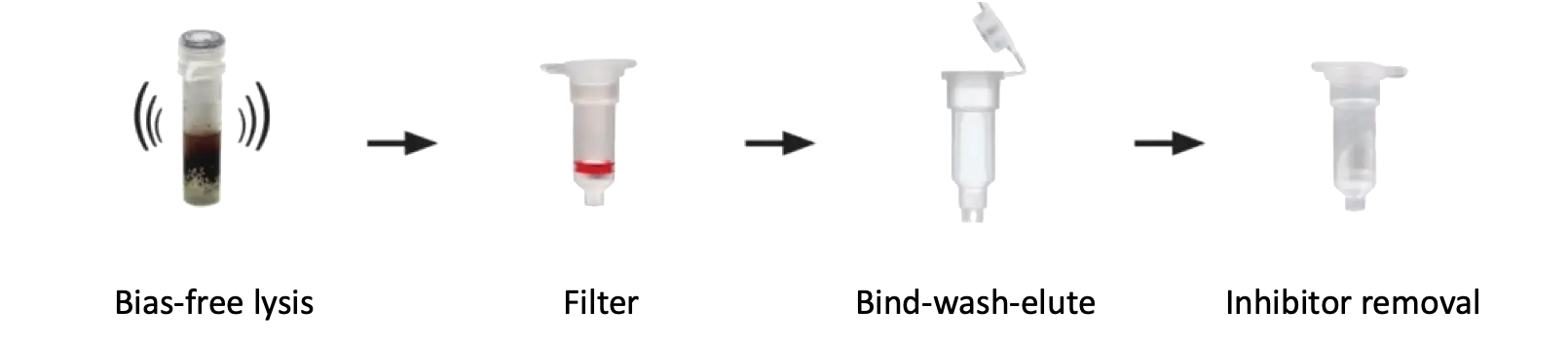
A wide array of samples can be easily loaded into the ZR BashingBead Lysis Tubes for complete and unbiased lysis, followed by filtering the lysate for debris removal. Samples are then processed using our patented Zymo-Spin Technology, making DNA extraction as easy as Bind, Wash, and Elute. Additionally, this kit is equipped with Zymo Research’s proprietary OneStep PCR Inhibitor Removal technology enabling PCR from the most PCR prohibitive environmental samples.
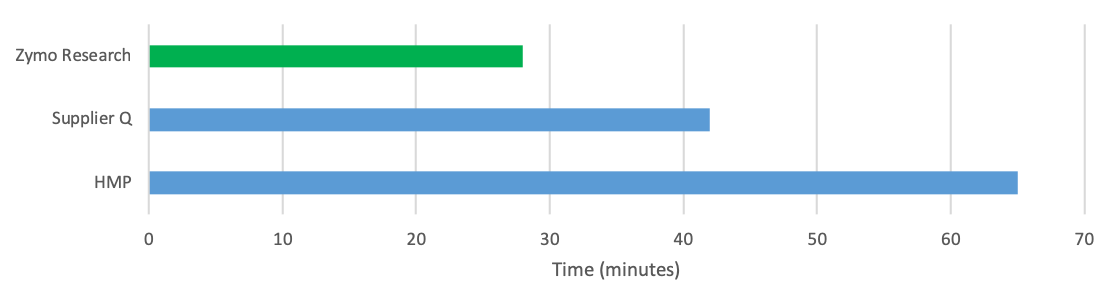
Sample to Pure DNA in <30 min
BUILT FOR ACCURACY
Unbiased Lysis of Samples
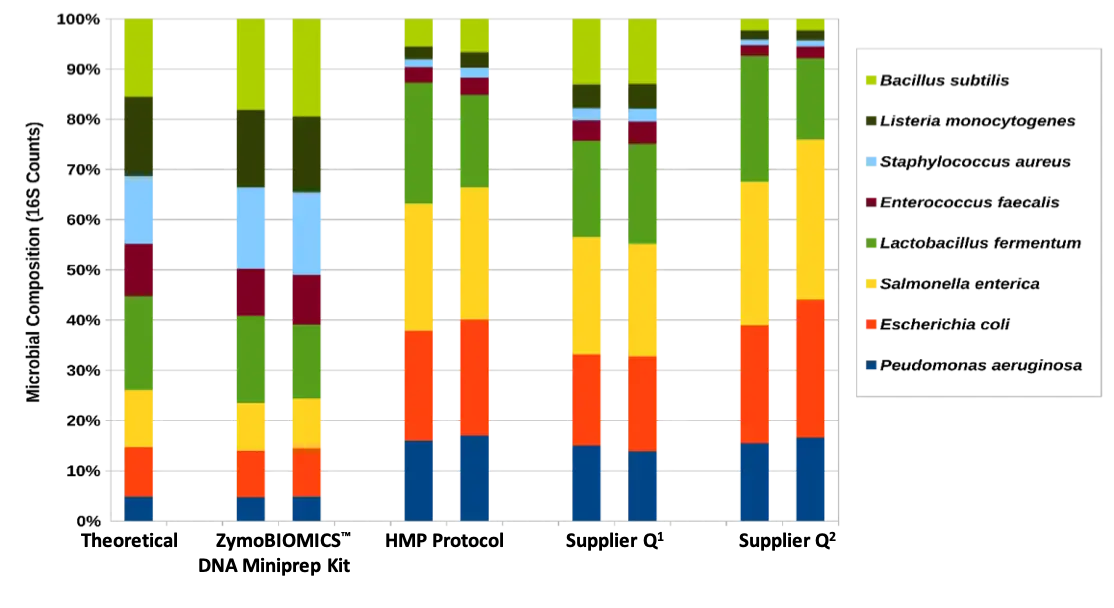
The ZymoBIOMICS DNA Miniprep Kit provides unbiased representation of the organisms extracted from the ZymoBIOMICS Microbial Community Standard. DNA was extracted from ZymoBIOMICS Microbial Community Standard using four different DNA extraction methods (ZymoBIOMICS DNA Miniprep Kit, Human Microbiome Project Protocol, Supplier Q1, and Supplier Q2) and analyzed using 16S rRNA gene sequencing.
Reveal the True Composition of Samples
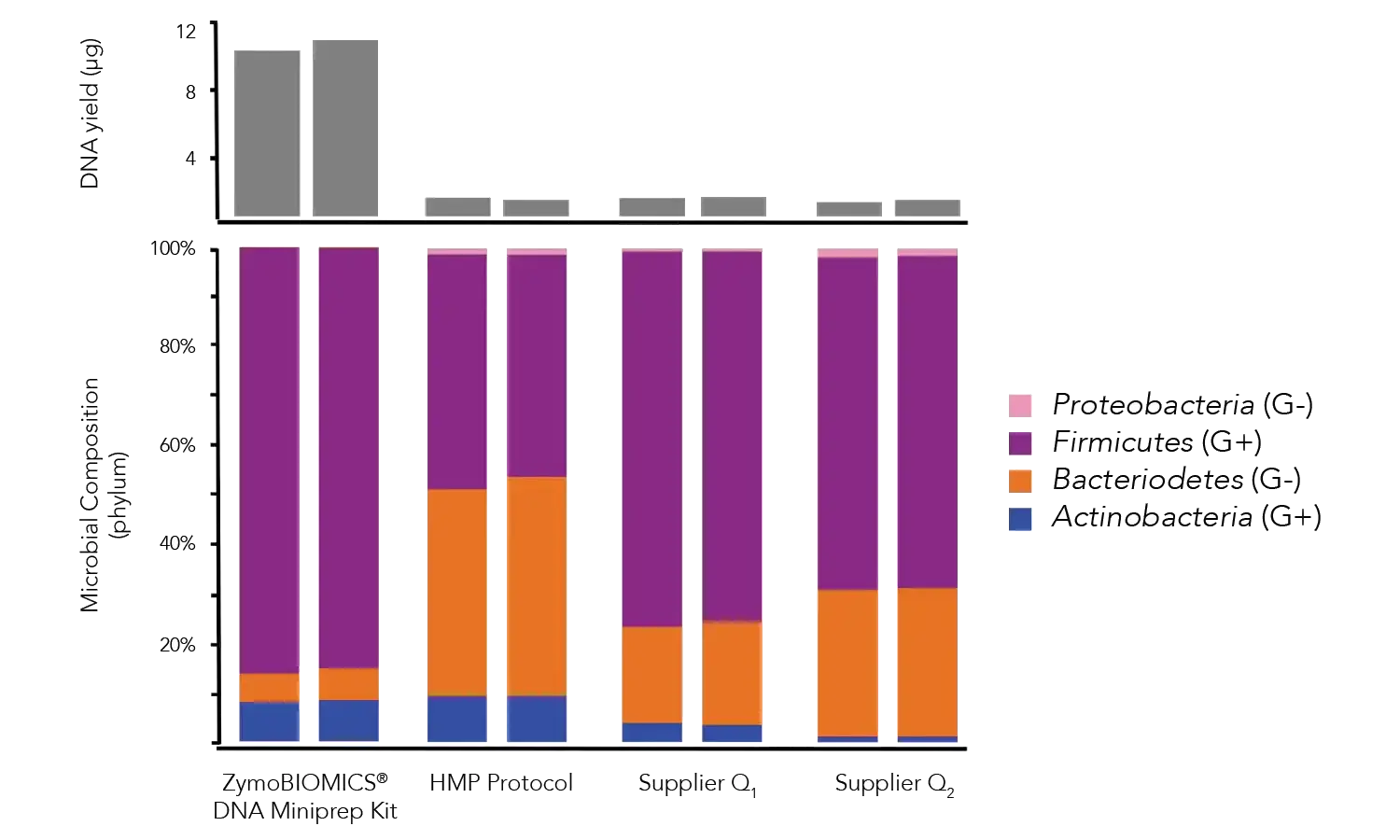
Total DNA from samples of human feces was extracted using the ZymoBIOMICS DNA Miniprep Kit, HMP fecal DNA extraction protocol, Supplier M and Supplier Q. The ZymoBIOMICS DNA Miniprep Kit yielded a higher total amount of DNA (upper). Additionally, the HMP, Supplier M and Supplier Q protocols over-represented easy-to-lyse microbes, e.g. Bacteroidetes and Proteobacteria, biasing the microbial profile.
Next-Gen sequencing ready

DNA extraction from soil samples using the ZymoBIOMICS DNA Miniprep Kit (D4300) and Qiagen DNeasy PowerSoil Pro Kit were analyzed with the Femto Bacterial DNA Quantification Kit (n=6). An additional set of eluates from Qiagen DNeasy PowerSoil Pro Kit extractions were cleaned up using the OneStep PCR Inhibitor Removal Kit (D6030) before analysis.
Kits shouldn't bring their own microbes.

Low Bioburden Certification
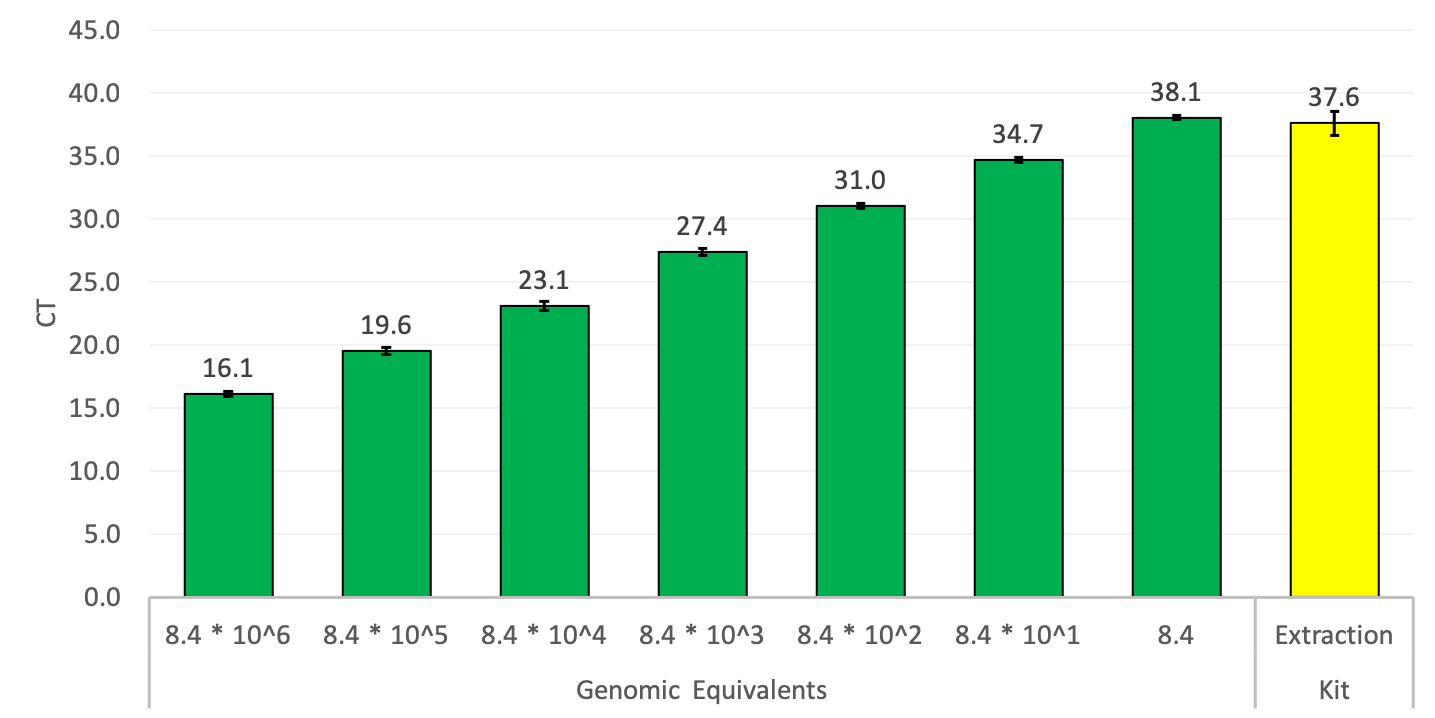
The bioburden of the kit was tested using real-time PCR with general bacterial primers. E.coli genomic DNA was used to create the standard curve.
Scale-up Analyses
Reproducible Sample Processing
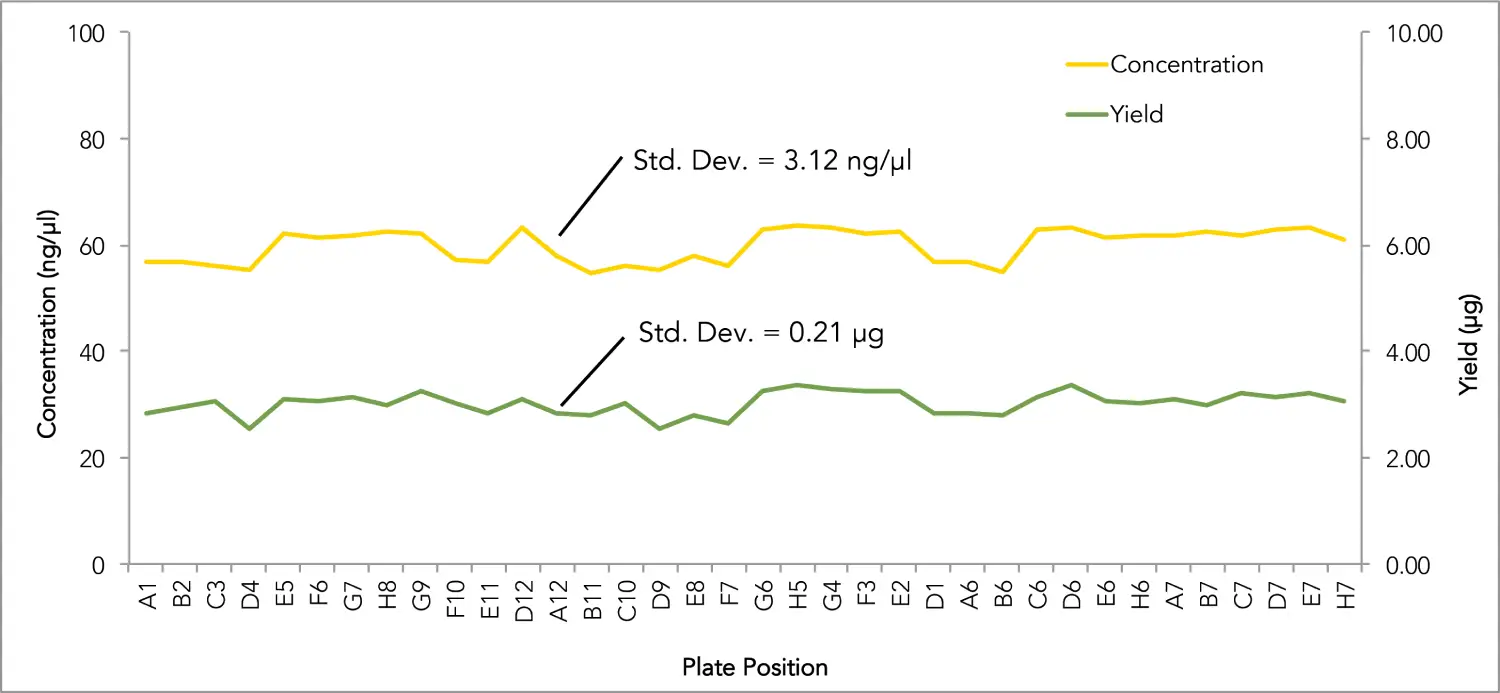
When 36 replicates of 200 µl blood samples from the same donor are processed, consistent and accurate yields are recovered in every sample. Average A260/230 ≥ 1.8.
No Cross-Contamination
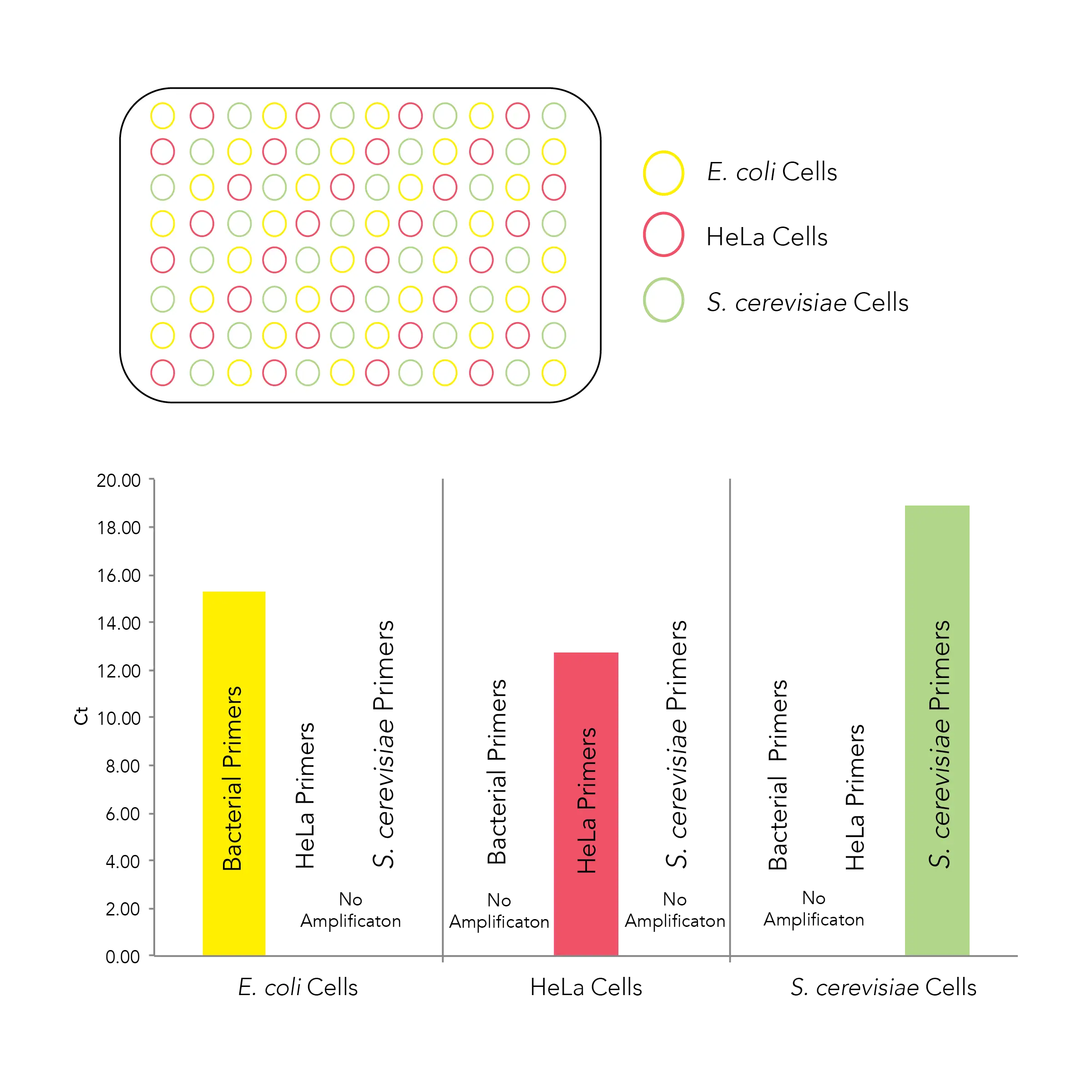
DNA is free from cross-contamination when purified with a liquid handler across a 96-well plate. The plate was set up with alternating rows of E. coli bacterial cells, HeLa cells, and S. cerevisiae yeast cells then simultaneously purified. Samples were evaluated using qPCR in duplicates.
| Product | Catalog # | Format | Binding Capacity | Protocol | Free Sample |
|---|---|---|---|---|---|
| Zymo |
D4300 | Spin Column | 25 µg |  |
Get Free Sample |
| Zymo |
D4301 | Spin Column | 5 µg |  |
|
| Zymo |
D4302 | 96 Magbead | 10 µg |  |
|
| Zymo |
D4309 | 96-Well Spin Column | 5 µg |  |
|
| Zymo |
R2002 | Spin Column | 100µg |  |
Get Free Sample |