Successfully Added to Cart
Customers also bought...
-
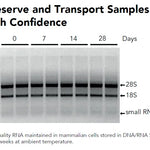 DNA/RNA Shield (50 ml)Cat#: R1100-50DNA/RNA Shield reagent is a DNA and RNA stabilization solution for nucleic acids in any biological sample. This DNA and RNA stabilization solution preserves the...
DNA/RNA Shield (50 ml)Cat#: R1100-50DNA/RNA Shield reagent is a DNA and RNA stabilization solution for nucleic acids in any biological sample. This DNA and RNA stabilization solution preserves the... -
 DNA/RNA Shield SafeCollect Swab Collection Kit (1 ml fill) (1 collection kit)Cat#: R1160The DNA/RNA Shield SafeCollect Swab Collection Kit is a user-friendly collection kit for stabilizing the nucleic acid content of samples collected with a swab. DNA/RNA...
DNA/RNA Shield SafeCollect Swab Collection Kit (1 ml fill) (1 collection kit)Cat#: R1160The DNA/RNA Shield SafeCollect Swab Collection Kit is a user-friendly collection kit for stabilizing the nucleic acid content of samples collected with a swab. DNA/RNA...
Highlights
- Sensitive and specific detection of 5-hydroxymethlycytosine (5-hmC) from a variety of samples
- Ideal for global 5-hmC quantitation and high throughput screening
- Streamlined workflow can be completed in 3 hrs
Original Manufacturer
Satisfaction 100% guaranteed, read Our Promise
Innovated in California, Made in the USA
Highlights
- Sensitive and specific detection of 5-hydroxymethlycytosine (5-hmC) from a variety of samples
- Ideal for global 5-hmC quantitation and high throughput screening
- Streamlined workflow can be completed in 3 hrs
Original Manufacturer
Satisfaction 100% guaranteed, read Our Promise
Innovated in California, Made in the USA
| Cat # | Name | Size | Price | |
|---|---|---|---|---|
| A4001-25 | Anti-5-Hydroxymethylcytosine Polyclonal Antibody | 25 µg | €107,90 | |
| D5425-5-1 | Control A (100 ng/µl) | 40 µl | €61,10 | |
| D5425-5-2 | Control B (100 ng/µl) | 40 µl | €61,10 | |
| D5425-5-3 | Control C (100 ng/µl) | 40 µl | €61,10 | |
| D5425-5-4 | Control D (100 ng/µl) | 40 µl | €61,10 | |
| D5425-5-5 | Control E (100 ng/µl) | 40 µl | €61,10 | |
| D5425-4-15 | HRP Developer | 15 ml | €13,30 | |
| D5425-1-15 | Coating Buffer | 15 ml | €32,60 | |
| D5425-2-30 | 10X ELISA Buffer | 30 ml | €36,70 | |
| D5425-3-100 | Anti-DNA HRP Antibody (100X) | 100 µl | €110,00 | |
| C2020 | 96-Well ELISA Plate | 12 x 8-Well Strips | €74,30 | |
Description
Performance
Technical Specifications
| Detection | ≥ 0.02% 5-hmC per 100 ng DNA |
|---|---|
| Equipment | Incubator and ELISA plate reader. A multi-channel pipettor is recommended. An automated plate washer may be used for blocking and wash steps. |
| Input | This protocol is optimized for 100 ng of input DNA/well. Compatible with DNA in the range of 20-200 ng/well. |
| Sample Source | Purified genomic DNA in water, Tris-EDTA, or similar. |
Resources
Documents
FAQ
We guarantee integrity of kit components for up to 6 months from date of purchase
Shipping of components is on blue and dry ice. Upon arrival, make sure to store buffers at 4°C, Control DNA and Anti-5hmC antibody at -20°C, and the Anti-DNA antibody can be stored at -20°C for 1 week (For long-term storage keep at -80°C).
The DNA ELISA controls are genomic DNA. Control A is E. coli genomic DNA as it comprises 0% 5-hmC. The other controls derive from various mouse tissues and the %5-hmC is specific to each tissue. The levels of 5-hmC are were quantify by mass-spec.
The optimal absorbance is at 405 nm, however 405-450 nm will work as well.
To calculate global %5-hmC, control DNAs and samples must be assayed on the same plate. Use the equation x=y-b/m and solve for x (%5-hmC).
1. Make sure the correct buffers were used to coat, block and wash the plate. 2. Double-check that HRP developer and the Anti-DNA HRP antibody were stored and prepared correctly.
1. Make sure that standard curve developed properly (R2 > 0.9). 2. Verify that all samples and controls are prepared correctly, mixed well before dilution and addition to the wells. 3. Ensure that anti-DNA HRP antibody was stored and prepared correctly before addition to the wells. 4. %5-hmC levels of your sample might be too low for the assay to detect (detection limit is 0.01% in 100 ng of denatured DNA).
Citations
Need help? Contact Us