Successfully Added to Cart
Customers also bought...
-
 DNA/RNA Shield (50 ml)Cat#: R1100-50DNA/RNA Shield reagent is a DNA and RNA stabilization solution for nucleic acids in any biological sample. This DNA and RNA stabilization solution preserves the...
DNA/RNA Shield (50 ml)Cat#: R1100-50DNA/RNA Shield reagent is a DNA and RNA stabilization solution for nucleic acids in any biological sample. This DNA and RNA stabilization solution preserves the... -
 DNA/RNA Shield SafeCollect Swab Collection Kit (1 ml fill) (1 collection kit)Cat#: R1160The DNA/RNA Shield SafeCollect Swab Collection Kit is a user-friendly collection kit for stabilizing the nucleic acid content of samples collected with a swab. DNA/RNA...
DNA/RNA Shield SafeCollect Swab Collection Kit (1 ml fill) (1 collection kit)Cat#: R1160The DNA/RNA Shield SafeCollect Swab Collection Kit is a user-friendly collection kit for stabilizing the nucleic acid content of samples collected with a swab. DNA/RNA...
Highlights
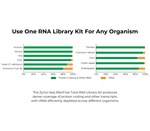
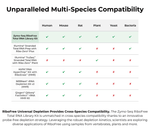
- Universal Depletion: Novel probe-free technology depletes rRNA from any organism.
- Simplest Library Prep: Simultaneous ligation of both adapters reduces hands-on processing.
- Automation Friendly: Streamlined protocol for increased scalability.
Original Manufacturer
Satisfaction 100% guaranteed, read Our Promise
Innovated in California, Made in the USA
Highlights
- Universal Depletion: Novel probe-free technology depletes rRNA from any organism.
- Simplest Library Prep: Simultaneous ligation of both adapters reduces hands-on processing.
- Automation Friendly: Streamlined protocol for increased scalability.
Original Manufacturer
Satisfaction 100% guaranteed, read Our Promise
Innovated in California, Made in the USA
This product was updated in 2023. Refer to FAQ-Q12 for details.
| Cat # | Name | Size | Price | Quantity |
|---|
| Cat # | Name | Size | Price | |
|---|---|---|---|---|
| R3001 | Zymo-Seq RiboFree Universal cDNA Kit | 12 Preps | €550,60 | |
| D3008 | Zymo-Seq UDI Primer Set (Indexes 1-12) | 12 indexes | €253,70 | |
| D3096 | Zymo-Seq UDI Primer Plate (Indexes 1-96) | 96 Indexes | €644,20 | |
| R3004-1-6 | Zymo-Seq Wash Buffer | 6 mL | €20,40 | |
| R3004-1-48 | Zymo-Seq Wash Buffer | 48 mL | €117,10 | |
| D3004-4-10 | DNA Elution Buffer | 10 ml | €17,40 | |
| D3004-4-50 | DNA Elution Buffer | 50 ml | €39,70 | |
| W1001-1 | DNase/RNase-Free Water | 1 ml | €12,30 | |
| W1001-4 | DNase/RNase-Free Water | 4 ml | €14,30 | |
| W1001-6 | DNase/RNase-Free Water | 6 ml | €18,40 | |
| W1001-10 | DNase/RNase-Free Water | 10 ml | €23,50 | |
| W1001-30 | DNase/RNase-Free Water | 30 ml | €26,50 | |
| 3DP-1002 | PCR Strip MagStand | 1 unit | €61,10 | |
Description
Performance
Technical Specifications
| Equipment Needed (user provided) | Thermal cycler with heated lid, magnetic stand for 0.2 mL PCR tubes, microcentrifuge for 0.2 mL PCR tubes and 1.5 mL microcentrifuge tubes, and a benchtop vortex mixer. |
|---|---|
| Input Quality | RNA should be free of DNA contamination and enzymatic inhibitors, with A260/A280 and A260/A230 ≥ 1.8. RNA with lower purity ratios (A260/A280 and A260/A230) should be treated with DNase I and purified with the RNA Clean & Concentrator™ (Cat. No. R1013) prior to processing. RNA should be suspended in water, TE, or a low-salt buffer.
|
| Library Storage | Libraries eluted in DNA Elution Buffer (provided) may be stored at ≤ 4°C overnight or ≤ -20°C for long-term storage. |
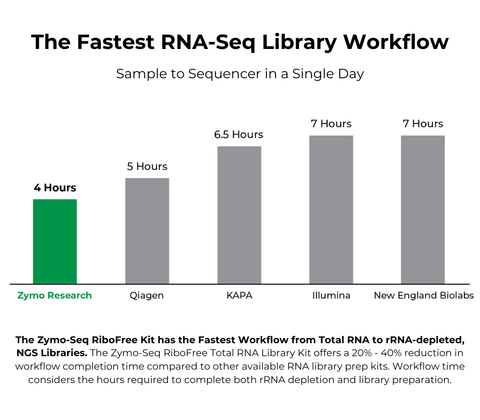
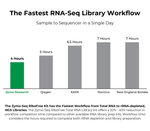
| Processing Time | As little as 4 hours |
| RNA Input | 10 – 250 ng of total RNA.
|
| Sample Input Material | RNA from any species |
| Sequencing Platform Compatibility | Libraries are compatible with all Illumina® sequencing platforms except HiSeq® X. Libraries are compatible with the Element AVITI™ system. While technically possible, Illumina® originally limits the applications on HiSeq® X exclusively for whole-genome libraries. Please confirm with the sequencing service provider for acceptability and additional details if expecting to sequence Zymo-Seq RiboFree Total RNA libraries on HiSeq® X series sequencers. |
Resources
Documents
FAQ
Purified total RNA from any organism. Please see our Multi-Organism Transcriptomics Application Note or find more product information here.
This kit is not designed for metatranscriptomics, and thus the performance cannot be guaranteed for such application. In our preliminary test using total RNA extracted from our ZymoBIOMICS Microbial Community Standard (D6300), using 100 ng as input and depleting for 2 hours yielded libraries where remaining rRNA reads were around 25%. Therefore, for rRNA depletion in metatranscriptomic samples with low complexity (<10 microbial species/strains), this kit could be suitable, and we recommend using a higher amount of total RNA as input whenever possible and optionally performing the Depletion Reaction for at least 2 hours. For further information, please contact tech@zymoresearch.com.
It is possible to prepare libraries using this kit with degraded RNA with low RIN scores, such as RNA from FFPE samples, although the performance may be negatively impacted. For optimal results using degraded samples, please refer to Appendix E in the kit protocol for detailed suggestions, or contact tech@zymoresearch.com.
DNA contamination will adversely impact the accuracy and sensitivity of quantitative measures of gene expression and differential gene expression analysis. Therefore, we recommend removing DNA contamination from the RNA input prior to performing the Zymo-Seq RiboFree workflow. For extracted RNA, we recommend using the RNA Clean & Concentrator-5 (R1013), which includes DNase I treatment and subsequent clean-up. This method has been validated for use with the Zymo-Seq RiboFree workflow.
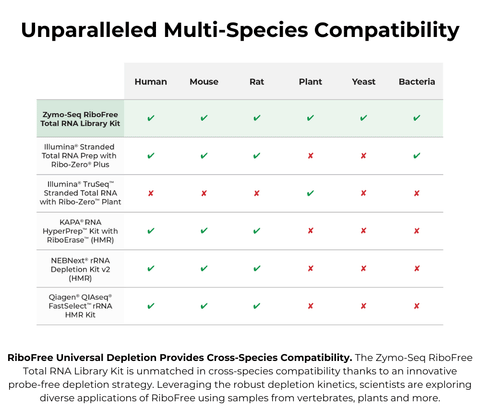
The Zymo-Seq RiboFree kit uses the input RNA as a template to drive the depletion of the reverse transcribed cDNA from the highly abundant sequences. This allows the depletion to be probe-free and universally compatible with RNA from any organism. Learn more about the probe-free depletion from this feature in Nature.
Generally, the random hexamer priming during reverse transcription and the reaction conditions during the depletion allows the production of appropriately sized inserts and final libraries for Illumina® sequencers.
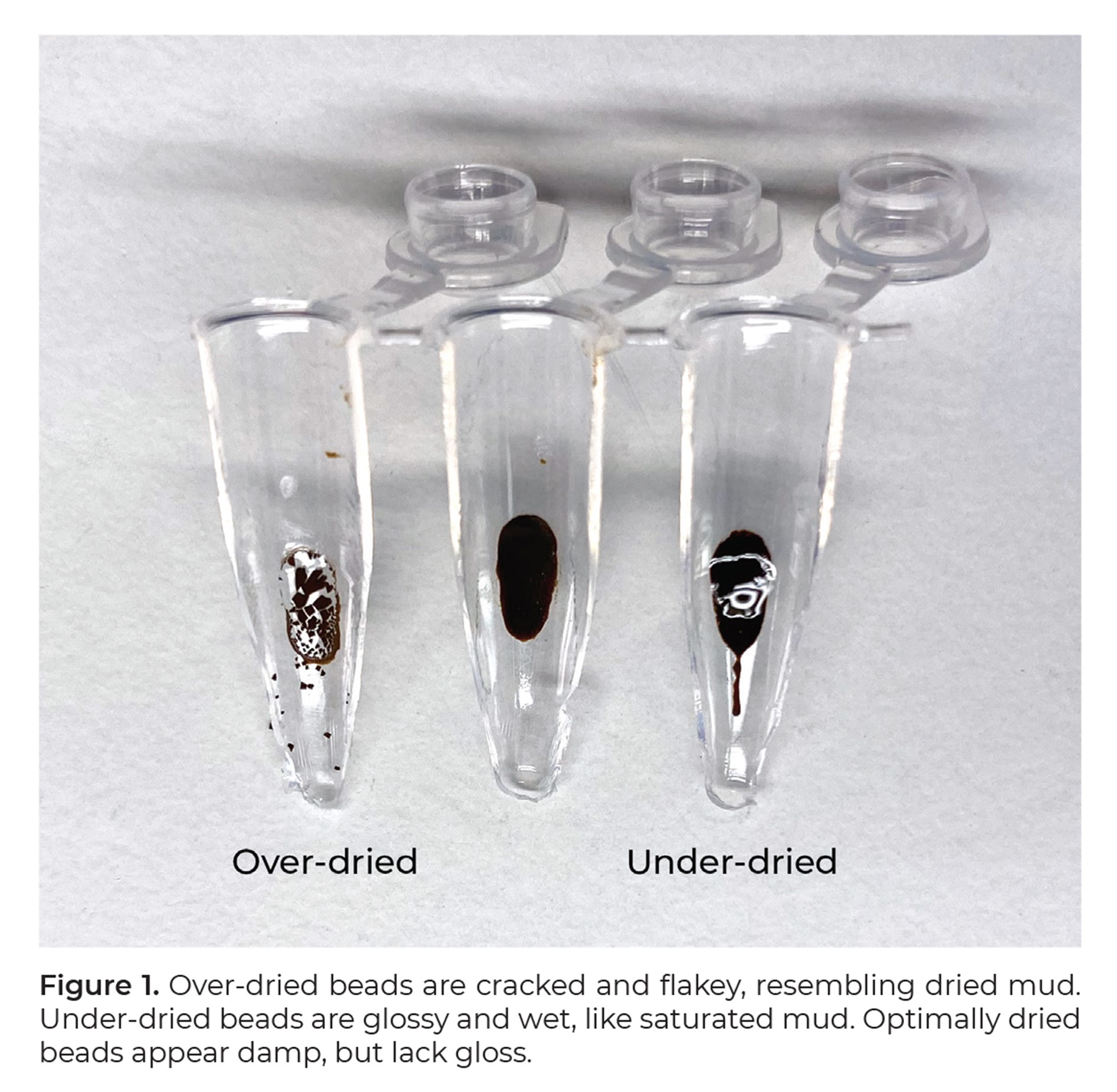
Humidity and temperature vary between labs. Therefore, while we don’t recommend a specific amount of time to wait, we do recommend adding the elution buffer to the bead pellet as soon as the wet gloss dries and leaves the pellet matte in appearance. By holding the tube up to a bright light, this change is more easily seen in real time.
Avoid adding the elution buffer when the bead pellet is still wet-looking, or after the pellet cracks as both may lead to a decrease in library yield. An example image showing different levels of “dryness” is here for your reference.

Besides being included in the 96-prep RiboFree kit (R3003), the UDI indexes 1-96 are also sold separately as the Zymo-Seq UDI Primer Plate (Cat. No. D3096).
For higher-plex applications requiring more than 96 unique dual indexes, we now offer the Zymo-Seq UDI Primer Plate (Indexes 1-96, 10-bp) and Zymo-Seq UDI Primer Plate (Indexes 97-192, 10-bp) (Cat. Nos. D3097 & D3098).
Each UDI primer set in the Zymo-Seq UDI Primer Set (Cat. No. D3008) can be used more than once as long as the libraries with the same UDI set ARE NOT pooled and sequenced on the same sequencing lane. For more information regarding the UDI primer sets, please refer to Appendix B in the kit protocol or contact tech@zymoresearch.com.
This kit is compatible with any Illumina® sequencer. It is not directly compatible with Ion Torrent®, Oxford Nanopore®, or PacBio® platforms.
Yes, the Zymo-Seq RiboFree depletion module is available as a stand-alone product, the Zymo-Seq RiboFree Universal cDNA Kit (R3001), and is compatible with library preparation kits that accept single-stranded DNA as an input.
There are many ways for you to be certain about which instruction manual to follow. One way is to check the cold reagent names: they are different between the two versions, and the updated version includes a reagent called "Depletion Reagent 4". Overall, compared to v1.3.0, the updated version has a lower minimum input (10 ng total RNA), a shorter depletion reaction time (1 hour for 100 ng of input), and one fewer round of bead cleanup. To learn more of the kit features since the update, or if you need an electronic copy of the previous instruction manual (v1.3.0), please contact tech@zymoresearch.com.
It is possible to prepare libraries with input lower than 10 ng, though the performance may be negatively impacted. For modifications of the protocol when using 1-9 ng of RNA as input, please refer to Appendix D in the kit protocol for detailed suggestions or contact tech@zymoresearch.com. We do NOT recommend extending the depletion reaction to run overnight.
Library yield can vary depending on factors such as the quality and quantity of the input RNA, and the cycle number of the PCR reaction for library amplification. In general, using 10 ng of Universal Human Reference RNA (RIN > 8.0) as input, the library yield is > 10 nmol/L when amplified with 12 PCR cycles and eluted to 20 µL of DNA Elution Buffer.
Yes, globin depletion may be achieved in addition to rRNA depletion with our Zymo-Seq RiboFree Total RNA Library Kit. The efficiency may vary based on factors such as the sample species, whole blood RNA integrity, globin transcript abundance, and input amount.
According to our internal testing with high-quality human whole blood total RNA, we observed <5% hemoglobin reads and <5% rRNA reads in the final libraries when using 100 ng of input, and <10% hemoglobin reads and < 10% rRNA reads when using 10 ng of input. Therefore, it is recommended to use higher input masses if lower globin is desired.
To learn more about how scientists applied the Zymo-Seq RiboFree Total RNA Library Kit in blood transcriptome profiling of multiple non-human species, please see this blog.
Product Video
Reviews
"This is probably the best RNA-seq library preparation kit I’ve used (and I have used many). It is very simple and gives clean, reproducible results at a very affordable price. I would highly recommend this product to anyone looking for a quicker and easier library preparation kit. Zymo has never let me down."
Galina
City of Hope
"I think this kit is faster and simpler to do than the Truseq RNA library prep kit. It was easy enough that I tried it once and got a nice library prep, with a high concentration and I did not make any mistakes in my first try! I will definitely buy this kit and recommend it to others without a doubt."
Rafael
UC Berkeley
"This RNA Library Kit was easy to use and recovered good efficiency."
Veronica L.
CICY Mexico
Citations
Need help? Contact Us