
One RNA Library Kit for ANY Organism
Zymo-Seq RiboFree® Total RNA Library Kit
Simplest Library Prep
Prepare ready-to-sequence libraries in as little as 4 hours
Suitable for FFPE
Only minimal modifications needed to the standard protocol
Automation Friendly
Streamlined protocol for increased scalability. Learn More.
Master Endless Applications with One Protocol
-
 Gene Expression Profiling
Gene Expression Profiling
-
 Microbial Transcriptomics
Microbial Transcriptomics
-
 Pathogen Detection
Pathogen Detection
-
 Plant Research
Plant Research
-
 Functional Annotation of Animal Genomes
Functional Annotation of Animal Genomes
-
 And Many More!
And Many More!
Achieve informative and efficient RNA-Seq analysis with our Zymo-Seq RiboFree® Total RNA Library Kit, featuring ribosomal RNA (rRNA) depletion to prevent it from overshadowing other RNA molecules. The Zymo-Seq RiboFree® Total RNA Library Kit integrates a unique universal depletion technology that enables rRNA depletion from the total RNA of any organism. Dive into the transcriptomes of less commonly studied organisms in addition to human, mouse, and rat samples, across various research applications.
Comprehensive Gene Detection for Every Transcriptome & Beyond
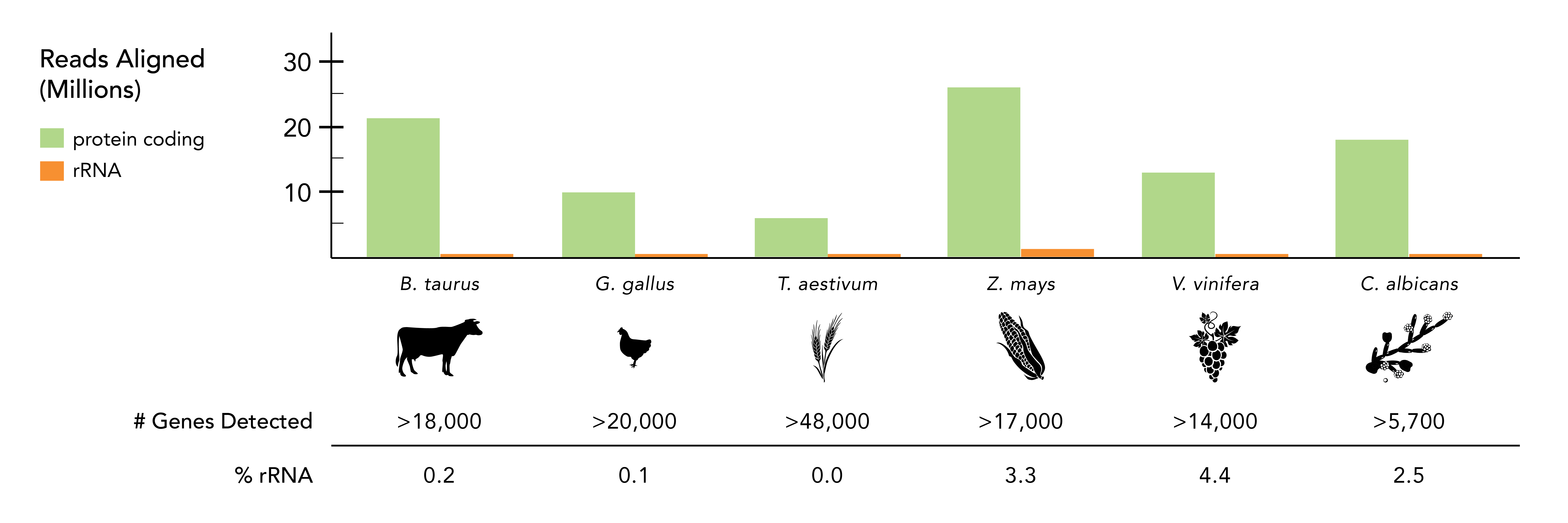
The Zymo-Seq RiboFree® Total RNA Library Kit Produces Dense Coverage of Protein Coding and Other Transcripts. The RiboFree® Universal rRNA Depletion technology boosted the number of reads for protein-coding and other non-rRNA transcripts to > 90% in samples across several kingdoms and phyla, detecting exceptional numbers of genes. Classification of the STAR-aligned reads was based on Ensembl annotations and RepeatMasker rRNA tracks from UCSC genome browser when applicable. “# Genes Detected” accounted for the number of unique gene IDs with TPM > 0.1.
The Fastest RNA-Seq Library Prep Workflow
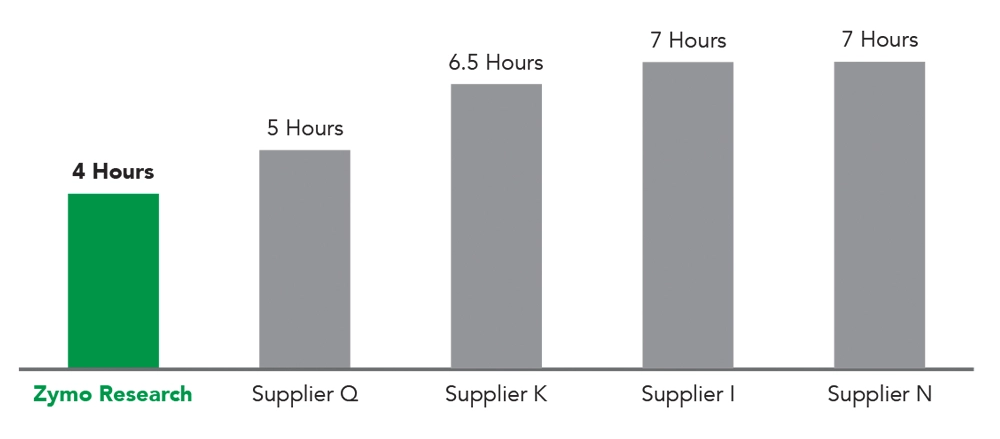
The Zymo-Seq RiboFree® Total RNA Library Kit has the Fastest Workflow from Total RNA to rRNA-depleted, NGS Libraries. The RiboFree® library prep workflow offers a 20% - 40% reduction in completion time compared to other similar RNA library prep kits. Workflow time considers the hours required to complete both rRNA depletion and library preparation.
Trusted by Scientists, Backed by Publications
The Zymo-Seq RiboFree® Total RNA Library Kit is underpinned by 150+ peer-reviewed studies, exemplifying its dependability and efficacy across varied applications. Here are select publications that highlight the breadth and effectiveness of our technology:
-
A single workflow for multi-species blood transcriptomics
 Read more
Read more
-
Choclo virus (CHOV) recovered from deep metatranscriptomics of archived frozen tissues in natural history biorepositories
 Read more
Read more
-
Individual bat virome analysis reveals co-infection and spillover among bats and virus zoonotic potential
 Read more
Read more
-
The rich non-coding RNA landscape of the Drosophila antennA
 Read more
Read more
-
Transcriptomes of an Array of Chicken Ovary, Intestinal, and Immune Cells and Tissues
 Read more
Read more
-
A photosynthesis operon in the chloroplast genome drives speciation in evening primroses
 Read more
Read more
Additional Resources
- Webinar: Watch our webinar on RNA-Seq applications.
- Blog: Read our latest articles on RNA sequencing advancements, and an overview of What is RNA-Seq

