Successfully Added to Cart
Customers also bought...
-
 DNA/RNA Shield (50 ml)Cat#: R1100-50DNA/RNA Shield reagent is a DNA and RNA stabilization solution for nucleic acids in any biological sample. This DNA and RNA stabilization solution preserves the...
DNA/RNA Shield (50 ml)Cat#: R1100-50DNA/RNA Shield reagent is a DNA and RNA stabilization solution for nucleic acids in any biological sample. This DNA and RNA stabilization solution preserves the... -
 DNA/RNA Shield SafeCollect Swab Collection Kit (1 ml fill) (1 collection kit)Cat#: R1160The DNA/RNA Shield SafeCollect Swab Collection Kit is a user-friendly collection kit for stabilizing the nucleic acid content of samples collected with a swab. DNA/RNA...
DNA/RNA Shield SafeCollect Swab Collection Kit (1 ml fill) (1 collection kit)Cat#: R1160The DNA/RNA Shield SafeCollect Swab Collection Kit is a user-friendly collection kit for stabilizing the nucleic acid content of samples collected with a swab. DNA/RNA...
Highlights
- Simple: Fast spin-column procedure yields pure yeast RNA without using glass beads or phenol.
- Versatile: Efficient RNA isolation from a broad spectrum of fungal species susceptible to Zymolyase.
- High-Quality: Isolated RNA is suitable for use in RT-PCR, Northern blotting, etc.
Original Manufacturer
100% satisfaction guaranteed, read Our Promise
Innovated in California, Made in the USA
Highlights
- Simple: Fast spin-column procedure yields pure yeast RNA without using glass beads or phenol.
- Versatile: Efficient RNA isolation from a broad spectrum of fungal species susceptible to Zymolyase.
- High-Quality: Isolated RNA is suitable for use in RT-PCR, Northern blotting, etc.
Original Manufacturer
100% satisfaction guaranteed, read Our Promise
Innovated in California, Made in the USA
| Cat # | Name | Size | Price | |
|---|---|---|---|---|
| W1001-6 | DNase/RNase-Free Water | 6 ml | €18,40 | |
| R1001-1 | YR Digestion Buffer | 3.2 ml | €28,50 | |
| R1001-2 | YR Lysis Buffer | 6.4 ml | €28,50 | |
| R1003-3-6 | RNA Wash Buffer | 6 ml | €22,40 | |
| C1001-50 | Collection Tubes | 50 Pack | €18,40 | |
| C1006-50-G | Zymo-Spin IIICG Columns | 50 Pack | €67,20 | |
| E1004 | Zymolyase | 1000 U | €107,20 | |
Description
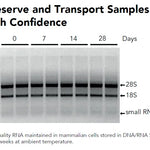
Performance
Technical Specifications
| Applicable For | Isolated RNA is well suited for downstream applications such as Northern blotting, RT-PCR, Next-Generation Sequencing, Microarray, hybridization and other sensitive applications |
|---|---|
| Elution Volume | ≥ 60 µl DNase/RNase-Free Water |
| Equipment | Microcentrifuge & Heat Block/Bath |
| Processing Time | 30 minutes |
| Processing Volume | ≤ 1.5 ml of Culture (1-5 x 107 cells) |
| Purity | Typical A260/A280 & A260/A230 ≥ 1.8 |
| Sample Source | Cell Culture (Colonies/Patches or Liquid Culture) |
| Size Range | ≥ 17 nt |
| Type | Total RNA |
| Yield | ≤ 25 µg of total RNA |
Resources
Documents
FAQ
Any strains susceptible to Zymolyase, which includes the following fungal genera: Asbya Kloekera Candida Kluyveromyces Debaryomyces Lipomyces Eremothecium Metschikowia Endomyces Pichia Hansenula Pullularia Hanseniaspora Saccharomyces Saccaromycodes Saccharomycopsis Schizosaccahromyces Torulopsis
Citations
Need help? Contact Us