Successfully Added to Cart
Customers also bought...
-
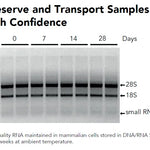 DNA/RNA Shield (50 ml)Cat#: R1100-50DNA/RNA Shield reagent is a DNA and RNA stabilization solution for nucleic acids in any biological sample. This DNA and RNA stabilization solution preserves the...
DNA/RNA Shield (50 ml)Cat#: R1100-50DNA/RNA Shield reagent is a DNA and RNA stabilization solution for nucleic acids in any biological sample. This DNA and RNA stabilization solution preserves the... -
 DNA/RNA Shield SafeCollect Swab Collection Kit (1 ml fill) (1 collection kit)Cat#: R1160The DNA/RNA Shield SafeCollect Swab Collection Kit is a user-friendly collection kit for stabilizing the nucleic acid content of samples collected with a swab. DNA/RNA...
DNA/RNA Shield SafeCollect Swab Collection Kit (1 ml fill) (1 collection kit)Cat#: R1160The DNA/RNA Shield SafeCollect Swab Collection Kit is a user-friendly collection kit for stabilizing the nucleic acid content of samples collected with a swab. DNA/RNA...
Highlights
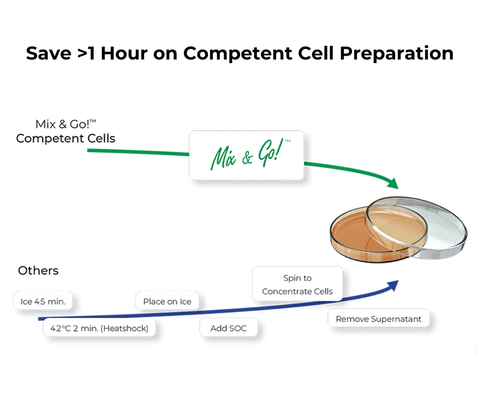
- Easy 3 Step Protocol: Produce reliable chemically competent E. coli in less than 45 minutes.
- Simple 20 sec Transformation: No heat shock! Just add DNA and spread.
- High Transformation Efficiencies: Achieve 108-109 transformants per µg of plasmid DNA.
Original Manufacturer
Satisfaction 100% guaranteed, read Our Promise
Innovated in California, Made in the USA
Highlights
- Easy 3 Step Protocol: Produce reliable chemically competent E. coli in less than 45 minutes.
- Simple 20 sec Transformation: No heat shock! Just add DNA and spread.
- High Transformation Efficiencies: Achieve 108-109 transformants per µg of plasmid DNA.
Original Manufacturer
Satisfaction 100% guaranteed, read Our Promise
Innovated in California, Made in the USA
| Cat # | Name | Size | Price | |
|---|---|---|---|---|
| T3001-2-10 | Mix & Go! 2X Stock Wash Buffer | 10 ml | €34,70 | |
| T3001-2-30 | Mix & Go! 2X Stock Wash Buffer | 30 ml | €64,20 | |
| T3001-3-10 | Mix & Go! 2X Stock Competent Buffer | 10 ml | €34,70 | |
| T3001-3-30 | Mix & Go! 2X Stock Competent Buffer | 30 ml | €64,20 | |
| T3001-4-20 | Mix and Go! Dilution Buffer | 20 ml | €14,30 | |
| T3001-4-60 | Mix and Go! Dilution Buffer | 60 ml | €31,60 | |
| M3015-100 | ZymoBroth | 100 ml | €38,20 | |
Description
Performance
Technical Specifications
| Compatible Strains | The Kit is optimized to work with K12 and B derivative strains. Some commonly used strains include, TOP10, ccdB survival, XL1 Blue and Stbl3. |
|---|---|
| Processing Time | ≤ 45 minutes |
| Product Storage | Please store the Wash Buffer (2X), Competent Buffer (2X), and Dilution Buffer at 0-8°C. The ZymoBroth can be stored at room temperature. |
| Transformation Efficiency | 108-109 transformants per µg of plasmid DNA for most common lab strains. |
Resources
Documents
FAQ
The Kit is optimized to work with K12 and B derivative strains. Some commonly used strains include, TOP10, ccdB survival, XL1 Blue and Stbl3. (https://blog.addgene.org/save-time-and-money-by-making-your-own-competent-cells) If working with a wild type strain of bacteria, these strains will generally have low transformation efficiencies. Therefore, we recommend trying our tips for boosting transformation efficiencies.
Prepare the overnight cultures in LB with shaking at 37°C. - If starting with already competent or electrocompetent cells, add 1-5 ul cells to 5ml of LB and shake overnight at 37°C. - If starting from fresh colonies, inoculate 5ml of LB with a single colony and shake overnight at 37°C.
The cells can be resuspended in 1X Wash Buffer and left on ice for up to 1 ½ hours without a significant decrease in transformation efficiency.
The cells can be resuspended in 1X Competent Buffer and left on ice for up to 1 ½ hours without a significant decrease in transformation efficiency.
1. Use ZymoBroth instead of SOB. 2. Expand the culture in ZymoBroth at lower temperatures (18-30°C, 25-26°C for practical reasons). Higher temperatures (i.e., 33-37 C) can decrease the transformation efficiency 2 to 10 fold. 3. Cells should be harvested early- to mid-log phase (OD600nm 0.4 – 0.6). Do not grow cells over 0.6 OD600nm as cells will be less competent as they reach late log or stationary phase. 4. Once the cells reach an OD between 0.4 and 0.6 and are placed on ice, make sure everything is ice cold during all steps of competent cell preparation. 5. Cool down the centrifuge to 0-4°C before use. 6. Cool down the following items at -20°C prior to use: - Sterile filter pipette tips - Sterile centrifuge bottles or conical tubes to be used for pelleting the cells. - Sterile microcentrifuge tubes to be used for aliquoting (can also be cooled down at -70°C prior to use) - A microcentrifuge tube rack (can also be cooled down at -70°C prior to use) 7. When resuspending the cells: - Do not remove the cells from the ice for too long - Make sure the tubes are in contact with the ice while resuspending - Only remove the cells from the ice for a couple of seconds to check on the progress. 8. Perform a Heat Shock/Outgrowth Procedure using the following protocol: 1. Incubate cells on ice for 5-10 min after addition of plasmid. 2. Incubate cells at 42°C for 45 seconds. 3. Add 450 ml of SOC to the cells. 4. Incubate at 37°C for 60 min with gentle shaking at 200-300 rpm. 5. Spread on a pre-warmed culture plate containing the appropriate antibiotic.
Yes, the buffers are very stable and can be stored at room temperature for a few months. Place your kit in the fridge (4°C) for longer storage. Please make sure the reagents are ice-cold before processing the harvested cells.
1. Aliquot the cells in the volume of transformation reaction that you wish to perform in order to minimize the number of manipulations. Repeated freeze/thaw cycles will cause some loss of competency. 2. Place the cooled microcentrifuge tube rack on top of ice (or dry ice), fill the rack with pre-chilled microcentrifuge tubes, aliquot the cells, and cap the tubes. Once you are finished, transfer the tubes to the -80°C for later use. Snap freezing in liquid nitrogen is not necessary.
Liquid nitrogen is only used to snap freeze the cells before storage. The most common method of storage is in a freezer kept below -70°C.
Transformation efficiency depends on several factors, including strain, growth conditions (e.g. growth medium, temperature, aeration, cell density at time of harvest), plasmid (e.g. size, origin of replication, resistance marker) and length of incubation with the plasmid. Therefore, efficiency cannot be guaranteed.
Reviews
"In molecular biology, bacterial transformation is a basic and essential step. Traditional methods of E. coli competent cell preparation take a lot of time and effort. This particular product from Zymo offers a simple method of competent cell preparation. This kit is very robust and the method is very simple. For the competent cells prepared by this method, heat shock is not required for the transformation. For transformation: thaw E. coli on ice and add required amount of DNA (1-5 ul) per 50 ul cells. Keep on ice for 5 minutes. Spread on a pre-warmed LB plate and incubate overnight at 37 deg C. The efficiency is near>10^7/µg (number of colonies observed after transformation). This kit is budget-friendly and at the same time, there is no compromise with the quality. Overall it's an efficient kit and highly recommended."
Yashavantha V.
"In our lab we do an average of 25 transformations per week, considering the high usage we prefer to prepare make our own cells. We always used the standard protocol to make the buffer which is always a pain especially as it just tends to be more laborious with the preparation of all the buffers. Here we just grow our bacteria at 18C until we reach the OD of 0.5 and then boom, just two simple steps with washing and resuspension and then the cells are ready. We did the same for DH5alpha and STBL3 strain and got competency in the range of 10^8."
Uday M.
"Very good, not expensive, easy to use, every lab can try this. Much cheaper than that of other companies, very fast, only takes 10 min."
Menemeng Z.
Citations
Need help? Contact Us