Successfully Added to Cart
Customers also bought...
-
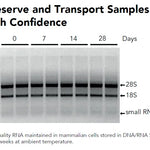 DNA/RNA Shield (50 ml)Cat#: R1100-50DNA/RNA Shield reagent is a DNA and RNA stabilization solution for nucleic acids in any biological sample. This DNA and RNA stabilization solution preserves the...
DNA/RNA Shield (50 ml)Cat#: R1100-50DNA/RNA Shield reagent is a DNA and RNA stabilization solution for nucleic acids in any biological sample. This DNA and RNA stabilization solution preserves the... -
 DNA/RNA Shield SafeCollect Swab Collection Kit (1 ml fill) (1 collection kit)Cat#: R1160The DNA/RNA Shield SafeCollect Swab Collection Kit is a user-friendly collection kit for stabilizing the nucleic acid content of samples collected with a swab. DNA/RNA...
DNA/RNA Shield SafeCollect Swab Collection Kit (1 ml fill) (1 collection kit)Cat#: R1160The DNA/RNA Shield SafeCollect Swab Collection Kit is a user-friendly collection kit for stabilizing the nucleic acid content of samples collected with a swab. DNA/RNA...
Highlights
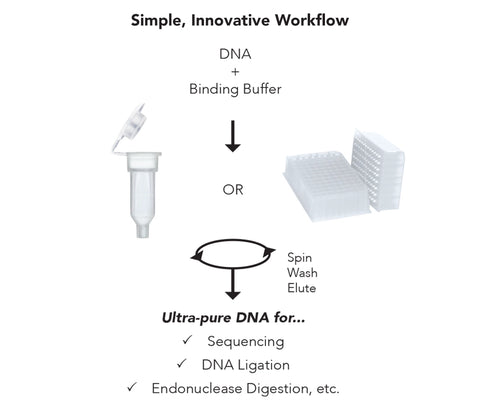
- Clean and concentrate up to 5 µg DNA with ≥ 6 µl elution in as little as 2 minutes with 0 µl wash residue carryover.
- Column design allows DNA to be eluted at high concentrations into minimal volumes of water or TE buffer.
- Eluted DNA is well suited for use in PCR, DNA sequencing, DNA ligation, endonuclease digestion, RNA transcription, radiolabeling, arrays, etc.
Original Manufacturer
Satisfaction 100% guaranteed, read Our Promise
Innovated in California, Made in the USA
Highlights
- Clean and concentrate up to 5 µg DNA with ≥ 6 µl elution in as little as 2 minutes with 0 µl wash residue carryover.
- Column design allows DNA to be eluted at high concentrations into minimal volumes of water or TE buffer.
- Eluted DNA is well suited for use in PCR, DNA sequencing, DNA ligation, endonuclease digestion, RNA transcription, radiolabeling, arrays, etc.
Original Manufacturer
Satisfaction 100% guaranteed, read Our Promise
Innovated in California, Made in the USA
| Cat # | Name | Size | Price | |
|---|---|---|---|---|
| D4003-2-24 | DNA Wash Buffer (Concentrate) | 24 ml | €40,80 | |
| D4003-1-25 | DNA Binding Buffer | 25 ml | €25,50 | |
| D4003-1-L | DNA Binding Buffer | 50 ml | €40,80 | |
| D4004-1-L | DNA Binding Buffer | 100 ml | €69,30 | |
| D4003-2-6 | DNA Wash Buffer (Concentrate) | 6 ml | €12,30 | |
| D3004-4-1 | DNA Elution Buffer | 1 ml | €13,30 | |
| D3004-4-4 | DNA Elution Buffer | 4 ml | €12,30 | |
| C1004-250 | Zymo-Spin IC Columns | 250 Pack | €278,90 | |
| C1003-50 | Zymo-Spin I Columns | 50 Pack | €61,10 | |
| C1003-250 | Zymo-Spin I Columns | 250 Pack | €261,60 | |
| C1001-1000 | Collection Tubes | 1000 Pack | €110,00 | |
| C1001-500 | Collection Tubes | 500 Pack | €64,20 | |
| C1004-50 | Zymo-Spin IC Columns | 50 Pack | €65,20 | |
| C1001-50 | Collection Tubes | 50 Pack | €18,40 | |
Description
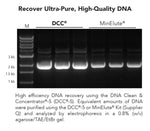
Performance
Technical Specifications
| Detergent Tolerance | ≤5% Triton X-100, ≤5% Tween-20, ≤5% Sarkosyl, ≤0.1% SDS |
|---|---|
| Elution Volume | ≥ 6 µl |
| Equipment | Microcentrifuge |
| Purity | A260/A280 > 1.8 |
| Sample Source | DNA from PCR, endonuclease digestions, DNA modification reactions, isotope/fluorescence labeling reactions, etc. |
| Size Range | 50 bp to 23 kb |
| Yield | ≤ 5 µg total DNA can be recovered. For DNA 50 bp to 10 kb the recovery is 70-95%. For DNA 11 kb to 23 kb the recovery is 50-70%. |
Resources
Documents
FAQ
The only difference is the cap. Everything else between the columns (max capacity, column matrix, etc.) is the same. Unsure which to use? It mainly comes down to preference. Some people like the capped columns for easy labeling or added security while others like the uncapped columns for faster sample processing.
Working with volumes below 50 µl can result in decreased recovery. Zymo Research recommend raising the starting volume to 100 µl with water to ensure optimal binding conditions.
Zymo Research recommends reloading columns no more than 5 times, as binding efficiency might decrease.
Oversaturation of the column can result in total DNA loss due to clogging of silica matrix.
Add an equal volume of ethanol (95-100%) to the sample and mix well. The sample is ready-to-bind and does not require DNA Binding Buffer. Proceed to Step 2.
Picogram levels of DNA can be recovered. The limitation is based on sensitivity of detection method.
The DNA will be eluted off the column during the wash step. Rebind samples using the appropriate amount of DNA Binding Buffer and wash the column with the properly prepared wash buffer.
We have a Zymoclean Gel DNA Recovery Kit that has chemistry compatible with agarose gel. However, if you have the DNA Clean & Concentrator kits on hand, you can clean up DNA from gel. You would need to purchase the Agarose Dissolving Buffer (Cat# D4001-1-50, D4001-1-100) separately, and follow the Zymoclean Gel DNA Recovery Kit protocol (link here).
Product Video
Reviews
"The elution volume is low enough to concentrate any sample very well and the efficiency is not compromised at all."
Shanshan L.
UT Southwestern
"I had low-volume, low-concentration, high-molecular weight genomic DNAs for high-throughput sequencing that I could not get to be perfectly clean without losing a significant amount of my (precious) sample. The Qiagen products were recommended by the company doing the sequencing, and they just ate up my sample. When I received this sample with my mini-prep kit, I was at the end of the line and figured I'd give it a try, despite being warned against anything involving spin columns that may shear the DNA. It worked beautifully. We can proceed with our project. No loss of sample, the gel looks the same (no sign of DNA shearing), and what an easy and fast protocol! Thank you, Zymo!"
Cindy T.
University of Iowa
"I sampled this kit next to a kit we already use. I had three of the same sample, two for the Zymo kit and one for the other. On one of the ones designated for your kit, I accidentally added the sample directly to the column filter, then added other reagents (I figured I might as well try it). The other I did as per the instructions. The one on which I goofed gave a similar purification and concentration as the other kit, while the one I did properly yielded approximately four times the DNA per µL as the other kit. I am very happy with this result, and when I rerun my samples, I plan to use your product."
Rosalind P.
University of Arkansas for Medical Sciences
Citations
Need help? Contact Us