Successfully Added to Cart
Customers also bought...
-
 DNA/RNA Shield (50 ml)Cat#: R1100-50DNA/RNA Shield reagent is a DNA and RNA stabilization solution for nucleic acids in any biological sample. This DNA and RNA stabilization solution preserves the...
DNA/RNA Shield (50 ml)Cat#: R1100-50DNA/RNA Shield reagent is a DNA and RNA stabilization solution for nucleic acids in any biological sample. This DNA and RNA stabilization solution preserves the... -
 DNA/RNA Shield SafeCollect Swab Collection Kit, 1ml (1 collection kit)Cat#: R1160The DNA/RNA Shield SafeCollect Swab Collection Kit is a user-friendly collection kit for stabilizing the nucleic acid content of samples collected with a swab. DNA/RNA Shield completely inactivates harmful pathogens...
DNA/RNA Shield SafeCollect Swab Collection Kit, 1ml (1 collection kit)Cat#: R1160The DNA/RNA Shield SafeCollect Swab Collection Kit is a user-friendly collection kit for stabilizing the nucleic acid content of samples collected with a swab. DNA/RNA Shield completely inactivates harmful pathogens...
EZ-96 DNA Methylation-Gold MagPrep
D5042 / D5043
EZ-96 DNA Methylation-Gold MagPrep
| Cat # | Name | Size | Price | Quantity |
|---|
Documents
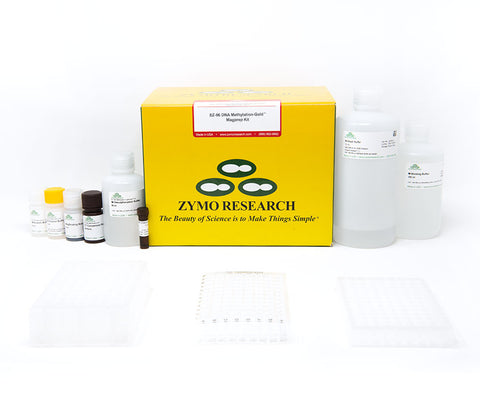
Highlights
- Complete, high-throughput bisulfite conversion of GC-rich DNA in less than 3 hours.
- A coupled heat denaturation/conversion reaction step streamlines the conversion of non-methylated cytosines to uracil.
- High throughput (96-well), automated desulphonation and recovery of bisulfite-treated DNA. Eluted, ultra-pure DNA is ideal for use in subsequent molecular-based analyses.
Description
| Applications | Purified, converted DNA is of high-quality and well-suited for downstream processes, including library preparation for Next-Generation sequencing, PCR amplification, etc. |
|---|---|
| Conversion | > 99% |
| Elution Volume | ≥ 25 µl |
| Equipment | Thermocycler with heated lid, heating element for 96-well plate, magnetic stand. |
| Input | 500 pg - 2 µg of DNA. |
| Processing Time | 3 hours |
| Recovery | > 70% |
| Sample Source | Purified genomic DNA, endonuclease-digested DNA, linearized plasmid DNA, etc. DNA should be high-quality and RNA-free. |
| Supplemental Info |
Q1: What leads to poor conversion efficiency/ low yields?
Poor conversion efficiency and low yields can be due to a variety of different experiment-specific conditions. Please contact Technical Support to discuss your specific experimental conditions and further troubleshoot with a product specialist.
Q2: How to quantify converted DNA?
For best results, keep the method of quantification consistent before and after bisulfite treatment:
- If quantifying with a NanoDrop, use dsDNA settings (50 μg/ml for Ab260 = 1.0) before treatment and use RNA settings (40 μg/ml for Ab260 = 1.0) after treatment.
- If quantifying with Qubit, use a dsDNA assay before treatment and use a ssDNA assay after treatment.
Q3: How to visualize converted DNA?
Following bisulfite treatment, DNA will be single stranded with limited non-specific base pairing at room temperature. To visualize, run the converted DNA on an agarose gel then chill the gel on ice or in an ice bath for 30 minutes. This will force enough base-pairing to allow intercalation of the ethidium bromide for the DNA to be visible. If using a Bioanalyzer or TapeStation instrument, use RNA kits and reagents to visualize the converted DNA.
Q4: What is the minimum DNA size that can be recovered?
> 50 bp.
Q5: How long is bisulfite converted DNA stable at -20 °C?
Converted DNA eluted in M-Elution Buffer can generally be stored at -20°C for 1-3 months. If longer term storage is necessary, we recommend storing at or below -70°C if possible. Bisulfite converted DNA is less stable than dsDNA; for best results, minimize freeze-thawing of converted DNA and use as soon as possible for downstream analysis.
Q6: Does bisulfite conversion only occur in a CpG context?
Bisulfite conversion will work regardless of context, so the kits are compatible with genomic DNA derived from plants and other species with high non-CpG methylation levels.
Q7: Is an incubation with desulphonation buffer for longer than 20 minutes recommended?
Leaving the desulphonation buffer on the column longer than recommended will cause more degradation and subsequently result in lower yields.
Q8: Which polymerase is recommended for amplification from bisulfite converted DNA?
ZymoTaq DNA Polymerase has been specifically designed for use in bisulfite amplification reactions. ZymoTaq is available as a stand-alone polymerase (E2001/E2002), PreMix (E2003/E2004), and qPCR PreMix (E2054/E2055).
Q9: Tips for bisulfite primer design?
| Cat # | Name | Size | Price | |
|---|---|---|---|---|
| D4100-5-8 | ||||
| D4100-5-16 | ||||
| D5041-6 | M-Elution Buffer | 40 ml | €37,80 | |
| D5003-1 | CT Conversion Reagent (96 Conversions) | 1 Bottle | €79,30 | |
| D5006-2 | M-Dilution Buffer-Gold | 7 ml | €12,70 | |
| D5007-6 | M-Elution Buffer | 8 ml | €12,70 | |
| D5040-4 | M-Wash Buffer | 72 ml | €85,50 | |
| D5040-5 | M-Desulphonation Buffer | 80 ml | €93,20 | |
| D5040-3 | M-Binding Buffer | 250 ml | €132,10 | |
| D5006-6 | M-Dissolving Buffer | 1.2 ml | €18,90 | |
| C2005 | Conversion Plate w/ Cover Foil | 2 Plates/Foils | €10,50 | |
| C2002 | Collection Plate | 2 Plates | €25,40 | |
| C2003 | Elution Plate | 2 Plates | €21,90 | |